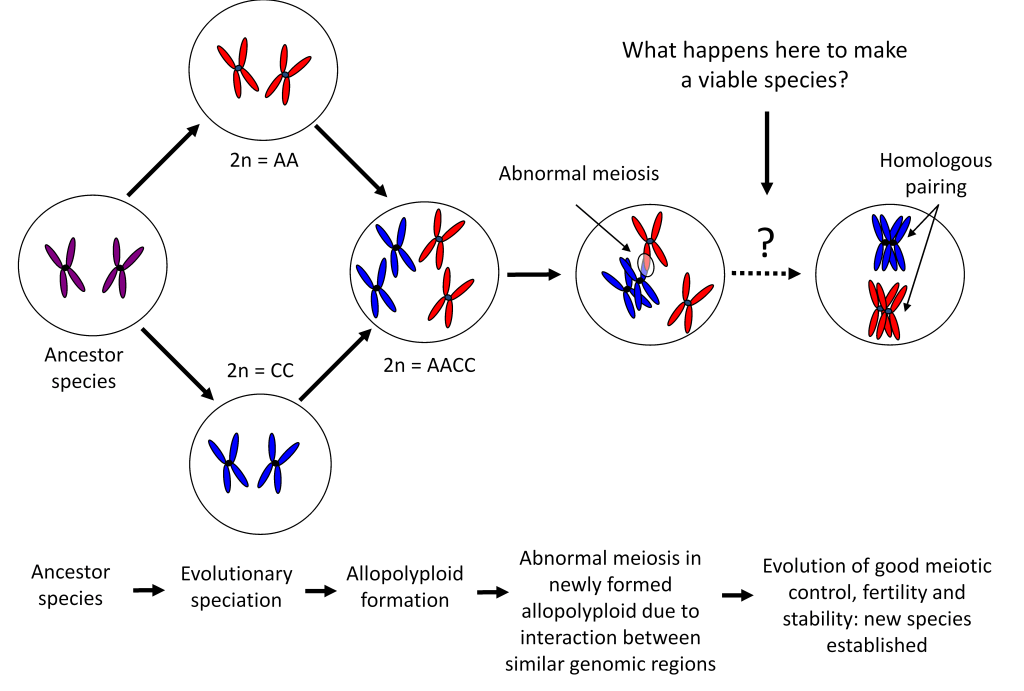
Allohexaploid Brassica
Although the Brassica genus contains both diploid (2n = 2x; one set of chromosomes/genome) and allotetraploid (2n = 4x; two sets of chromosomes/genomes) species, no naturally occurring three-genome allohexaploid exists. We aim to synthesise novel allohexaploid Brassica genotypes and investigate genome stability and fertility in these lines. A new allohexaploid Brassica crop will hopefully demonstrate improved hybrid vigour and adaptability, allowing incorporation of useful traits from all six cultivated Brassica diploid and allotetraploid species.

Hybrid speciation

Presence or absence of additional chromosomes (aneuploidy) is a phenomenon found to be increasingly common in nature. We are interested in whether aneuploidy can lead to speciation, or at least formation of new, stable karyotypes in Brassica. Chromosome and allele inheritance in different populations of novel interspecific hybrid types are being tracked across generations to determine what role aneuploidy may play in hybrid speciation in Brassica, or if new, stable genomes can be established over time.

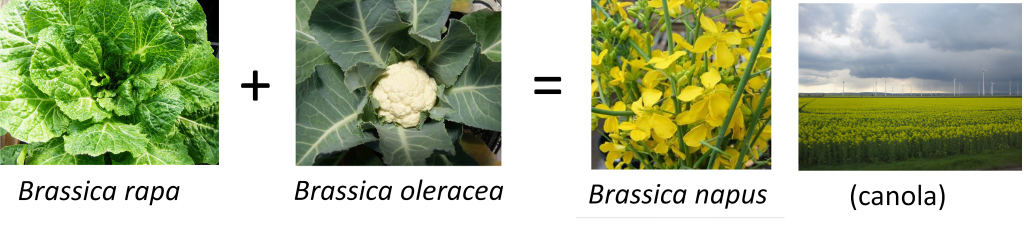
Recreating genomically stable rapeseed
In order to increase genetic diversity in highly inbred crop rapeseed (Brassica napus), a common method is to “recreate” this species by making new hybrids between rapeseed progenitor species B. rapa and B. oleracea. However, these hybrids also have unstable genomes due to poor control of meiosis, and lose chromosomes, and hence essential genetic information for plant growth and fertility, from generation to generation. The reason for this genome instability is unknown, particularly since “natural” B. napus is genomically stable. We aim to investigate genomic stability in a large set of human-made hybrid rapeseed genotypes using high-throughput marker genotyping, fertility phenotyping and cytogenetics. Identification of the mechanism/s of genomic stability in B. napus will not only provide fascinating insights into the evolutionary history of this species, but will be immediately useful for informing and assisting in transfer of useful genetic diversity into rapeseed.

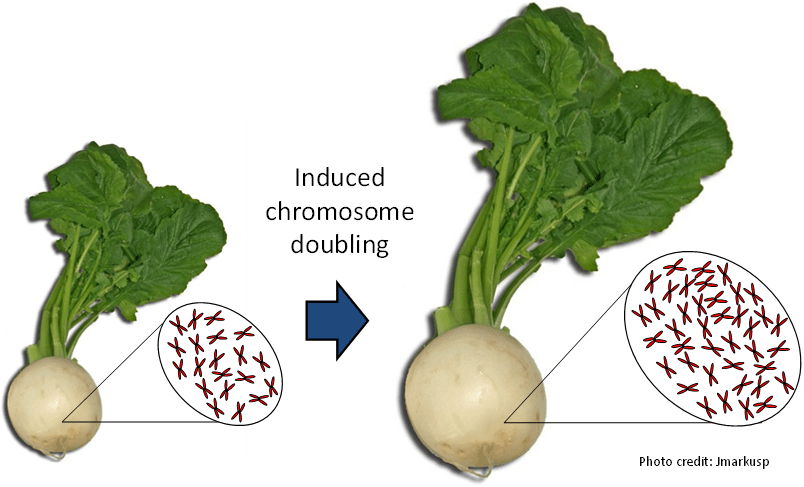
Towards autopolyploid Brassica crops
Polyploids often have larger cell and organ sizes than diploids. Hence, induced polyploidy, where chromosome numbers within a species are artificially doubled, has great potential for plant breeding, particularly of root and vegetable crops. In turnip (diploid Brassica rapa) and related vegetable species, targeted breeding efforts in the 1970s and 80s resulted in the successful release of a number of tetraploid crop types. However, these polyploid induction breeding strategies were hindered by problems such as poor fertility and vigour, and were subsequently abandoned. We propose to use modern genotyping, sequencing and cytogenetics technologies to identify the factors responsible for success and failure of induced polyploidy breeding in these crops. Such hybrids have immense potential for yield improvement as a result of increased heterosis, if stable tetraploid crop types can be created on demand.
Introgressing blackleg resistance from black mustard into rapeseed
Crop wild relative/minor crop species black mustard (B. nigra) is known to carry novel several sources of resistance to major rapeseed (B. napus) crop pathogen blackleg (Leptosphaeria maculans, convar Phoma lingam). We aim to identify these novel resistance sources and introgress the genetic loci responsible into a rapeseed background, providing useful germplasm with blackleg disease resistance for further breeding efforts.
